Quantitative description and classification of protein structures by a novel robust amino acid network: interaction selective network (ISN)
Abstract
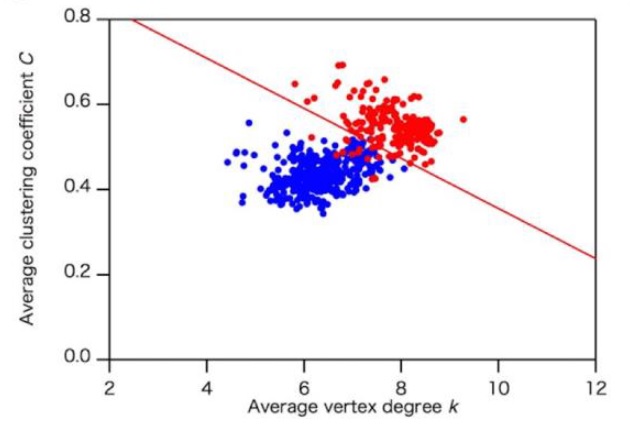
To quantitatively categorize protein structures, we developed a quantitative coarse-grained model of protein structures with a novel amino acid network, the interaction selective network (ISN), characterized by the links based on interactions in both the main and side chains. We found that the ISN is a novel robust network model to show the higher classification probability in the ……
Read more on Scientific Reports
Article information:
Konno, S., Namiki, T., and Ishimori, K.,
Quantitative description and classification of protein structures by a novel robust amino acid network: interaction selective network (ISN). Sci Rep 9, 16654 (2019).
DOI: 10.1038/s41598-019-52766-6